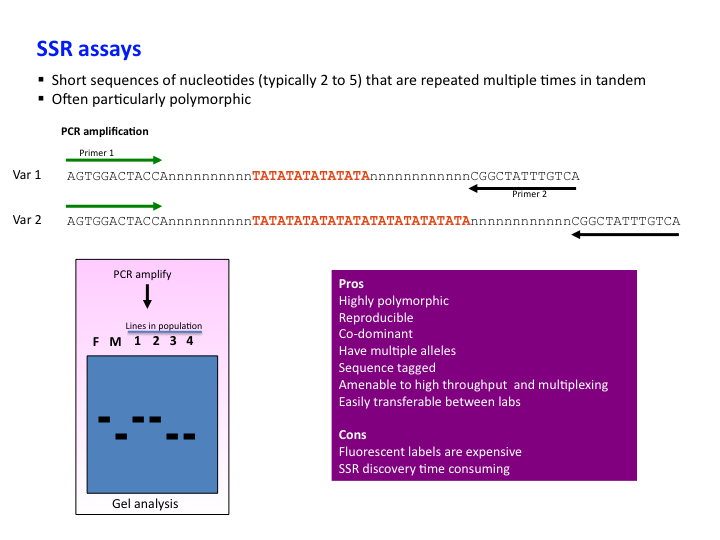
Identification and Sequence-Based Validation of the EST-SSR Markers from Calotropis procera | SpringerLink
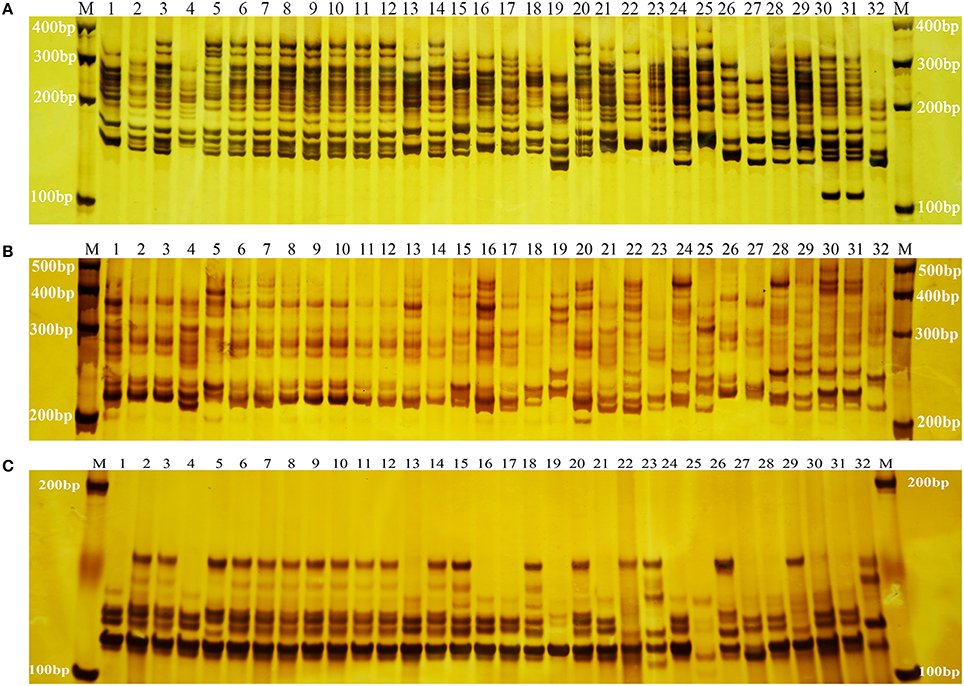
Frontiers | Development of SSR Markers and Assessment of Genetic Diversity in Medicinal Chrysanthemum morifolium Cultivars
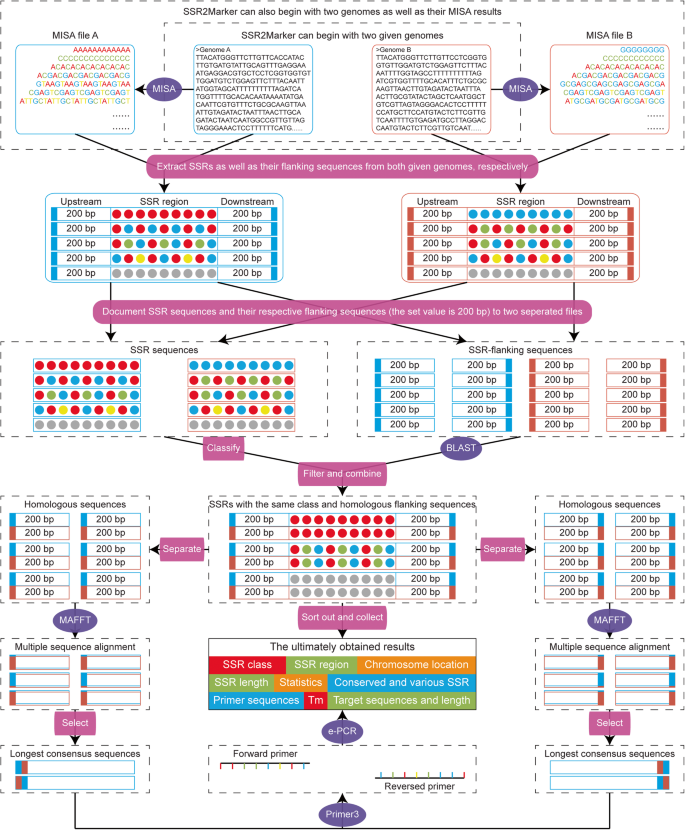
SSR2Marker: an integrated pipeline for identification of SSR markers within any two given genome-scale sequences | Molecular Horticulture | Full Text

Comprehensive functional analysis and mapping of SSR markers in the chickpea genome (Cicer arietinum L.) - ScienceDirect
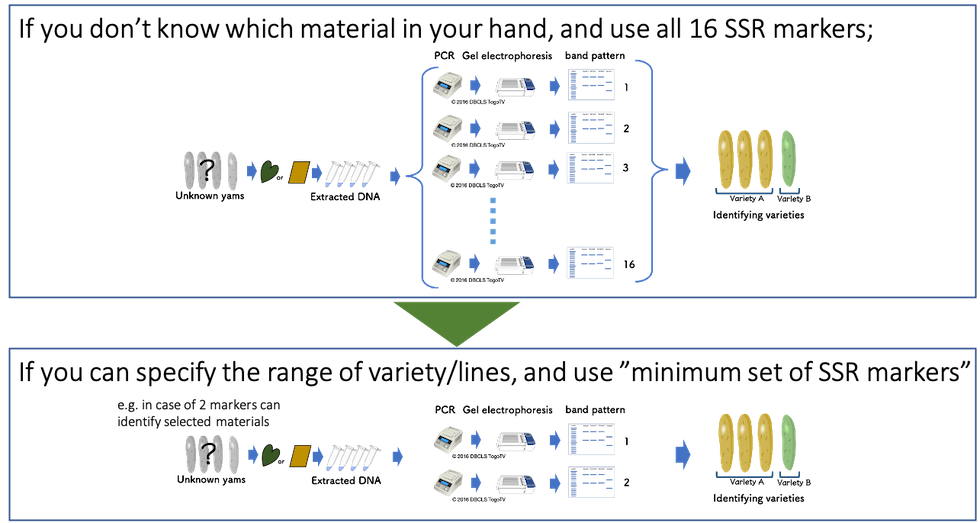
Yam Variety Identification Toolkit:Step 1 | Japan International Research Center for Agricultural Sciences | JIRCAS
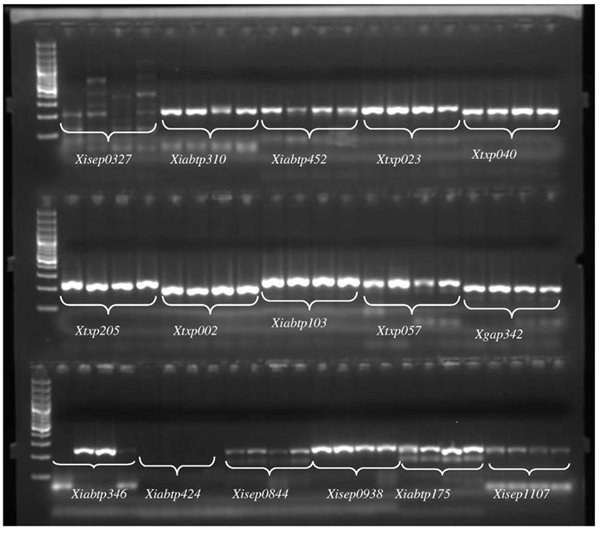
![Frontiers | New Hypervariable SSR Markers for Diversity Analysis, Hybrid Purity Testing and Trait Mapping in Pigeonpea [Cajanus cajan (L.) Millspaugh] Frontiers | New Hypervariable SSR Markers for Diversity Analysis, Hybrid Purity Testing and Trait Mapping in Pigeonpea [Cajanus cajan (L.) Millspaugh]](https://www.frontiersin.org/files/Articles/243873/fpls-08-00377-HTML-r1/image_m/fpls-08-00377-g001.jpg)
Frontiers | New Hypervariable SSR Markers for Diversity Analysis, Hybrid Purity Testing and Trait Mapping in Pigeonpea [Cajanus cajan (L.) Millspaugh]
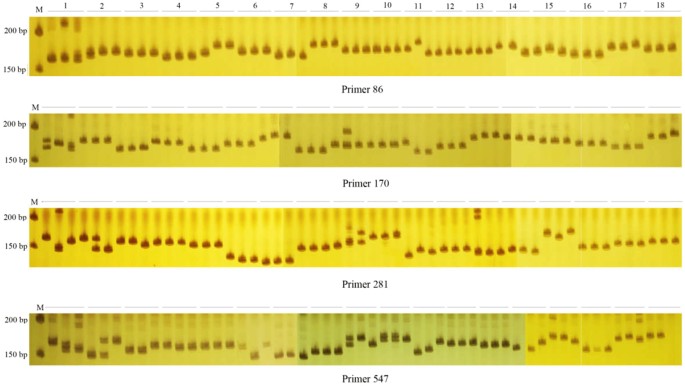

Cross-species transferability of EST-SSR markers developed from the transcriptome of Melilotus and their application to population genetics research | Scientific Reports

A general protocol for developing SSR markers with a SSR-enrichment step. | Download Scientific Diagram
Development of novel EST-SSR markers for ploidy identification based on de novo transcriptome assembly for Misgurnus anguillicaudatus | PLOS ONE

Genome-wide simple sequence repeats (SSR) markers discovered from whole-genome sequence comparisons of multiple spinach accessions | Scientific Reports
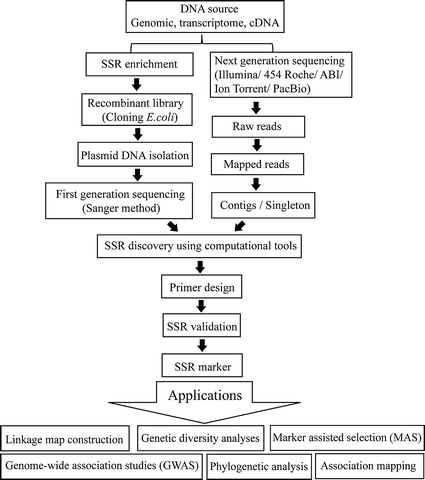











![PDF] Genic microsatellite markers in plants: features and applications. | Semantic Scholar PDF] Genic microsatellite markers in plants: features and applications. | Semantic Scholar](https://d3i71xaburhd42.cloudfront.net/441709d225d99041ac054dd81d8a130455e8a18b/2-Figure1-1.png)